Note: This tutorial assumes that the computer to be used already has a functioning installation of the latest version of the Starlink software suite (the latest release is available for download here).
You must initialise the Starlink software following the instructions on the release page in the same terminal you are going to run this tutorial in.
Basic Pipeline Reduction
This tutorial demonstrates how to run a standard, basic reduction of a SCUBA-2 dataset using the ORAC-DR pipeline software using different data reduction recipes and provides an introduction to basic Gaia use.
- Obtain a copy of the file JCMT_SCUBA-2_tutorial1_2016.tar.gz containing the raw data from here (file size: ~650 MB). This file contains SCUBA-2 850-micron band observational data for G34.3+0.2.
Copy this file to a suitable directory to work in (e.g. your local home directory, ~/ , in the following example):
cp JCMT_SCUBA-2_tutorial1_2016.tar.gz ~/ - Switch to the working directory, open the tarball and change to the newly-created directory containing the raw data:
cd ~/
tar xvzf JCMT_SCUBA-2_tutorial1_2016.tar.gz
cd tutorial/raw/This directory should contain a number of .sdf files and a README document. This can be checked by listing the contents of the directory by typing
ls - Create an index file (called mylist) listing all of the input files, with explicit pathnames, and move it to the reduced directory, and switch to that as the current working directory (ORAC-DR will be run from there later on):
ls -d -1 $PWD/*.sdf > mylist
mv mylist ../reduced/
cd ../reduced/ - Set up ORAC-DR for SCUBA-2 850-micron data processing, and specify that the reduced files will be written to the current working directory:
oracdr_scuba2_850 -cwdThe -cwd option specifies that the current working directory should be used. Depending on your environment variable settings, it may not be needed.
By default, ORAC-DR will issue a warning that the default input data directory does not exist. It is therefore necessary to manually set the ORAC_DATA_IN environment variable. In case there is uncertainty as to which shell is currently in use, type the following to find out the answer:
echo "$SHELL"If in the Bash or zsh shell, type:
export ORAC_DATA_IN=~/tutorial/raw/Or, if in the TC-shell or C-shell, type:
setenv ORAC_DATA_IN ~/tutorial/raw/ - It should now be possible to run ORAC-DR by typing the following:
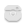
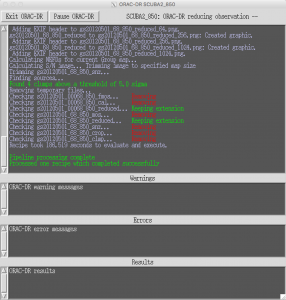
oracdr -loop file -files mylistThis should launch a separate window showing the progress of the data reduction. It will also indicate any problems encountered, and report as files are created and/or deleted.
Once the reduction process has completed, press the Exit ORAC-DR button to close the window. A number of files should have been produced as a result of the reduction process, including various logs, NDF and PNG preview image files.
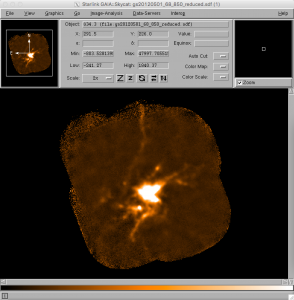
- It is also possible to examine the reduced group dataset in Gaia. This can be done by typing:
gaia gs20120501_68_850_reduced.sdf &This should launch a main Gaia window with the reduced image in it. To get a better sense of the more extended structure in the source, use the View drop-down menu, select Cut Levels and use the Auto Set option of 98%. It is also trivial to zoom in and out of the image using the two Z buttons.
Gaia has many other features to aid in image analysis. Feel free to experiment!
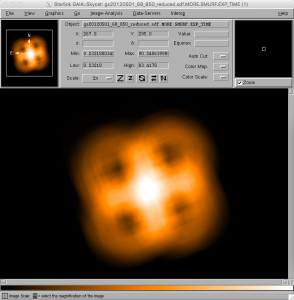
- As well as examining the reduced image data in Gaia, it is also possible to view other components of the NDF file. For example, the dataset also includes maps of the weights used at each position, the exposure times, and so on. In order to view these, the general strategy is to locate the supplementary Gaia window with the title Select NDF in container file, then select the desired table entry, and finally select one of the buttons near the bottom of that window. For example, to view the variance map of the science data, click on the top row of the table (number 1) to choose the science dataset, and then press the Variance button to see the science data variance map in the main Gaia window. Similarly, to view the exposure time map for the field, select the third row of the table (number 3 / EXP_TIME) and click on the data button. It should be noted that the levels of each of these maps can be very different, so it may be necessary to modify the cut levels again, as in step 6.
- Each SCUBA-2 science observation specifies a default data reduction recipe in the header information. This is originally set by selecting a recipe option in the Observing Tool during MSB preparation. To view the default recipe for a given dataset, switch to the raw data directory:
cd ../raw/If you set up the KAPPA package by typing
kappa
And seeing in your terminal
KAPPA commands are now available -- (Version 2.6-4)Type kaphelp for help on KAPPA commands.
Type ‘showme sun95’ to browse the hypertext documentation.See the ‘Release Notes’ section of SUN/95 for details of the
changes made for this release.You can use either of the following commands to examine the relevant part of the FITS header component of the data:
fitsval s8a20120501_00068_0001.sdf RECIPEor
fitslist s8a20120501_00068_0001.sdf | grep RECIPEThe default recipe for the example dataset is REDUCE_SCAN.
- It is also possible to re-run the pipeline reduction process using a different recipe from the default specified in the FITS header by simply appending the name of the recipe to the command line instruction. To try this, switch to the reduced data directory and try re-running the pipeline reduction with a different recipe:
cd ../reduced/
oracdr -loop file -files mylist REDUCE_SCAN_EXTENDED_SOURCESNote that when this is run, ORAC-DR will issue a warning about the suitability of the specified recipe for the DAISY mode data used in this tutorial. This demonstrates that care should be taken when manually specifying recipes.
To regenerate the original default reduction, just re-run ORAC-DR with the default recipe as before:
oracdr -loop file -files mylist
Other JCMT data reduction/analysis tutorials are available here.