Note: This tutorial assumes that the computer to be used already has a functioning installation of a recent (2016A or newer) version of the Starlink software suite (the latest release is available for download here). The user should download the relevant data for this tutorial:
- Dataset for SCUBA-2 Data Reduction/Analysis Tutorials 1 – 3 & 5 (Filesize: ~650 MB)
- Dataset for Heterodyne Data Reduction/Analysis Tutorial 1 also required for SCUBA-2 Data Reduction/Analysis Tutorial 5 (Filesize: ~77MB)
The user is also advised to first obtain basic familiarity with SCUBA-2 datasets, via the relevant JCMT SCUBA-2 web Tutorials 1 and 2 and heterodyne data reduction, via the relevant JCMT heterodyne web Tutorial 1 (specifically using data: 20070705 Scans 38+39).
CO (3-2) contamination estimation
It has long been reported in the literature that the CO (3-2) molecule provides additional emission at 850 micron than the emission produced by dust alone. To fully appreciate this tutorial you are encouraged to read: Molecular line contamination in the SCUBA-2 450 and 850 μm continuum data by Drabek et al. 2012, Parsons et al. 2018 and also the SCUBA-2 contamination page.
Before proceeding further you will need to ensure the Starlink software is loaded in your terminal, and you have loaded the analysis tools package; kappa:
starlink
kappa
if you have issues please ensure you have installed starlink correctly, you can also see the Starlink support mailing list for questions/concerns (or email helpdesk@eaobservatory.org).
STEP 1: Creating a HARP reference input file
If you have run the Heterodyne Instrument Data Reduction/Analysis Tutorial 1 on observations 20070705 38 and 39 and have produced file: ga20070705_38_1_integ.sdf you can skip this first step
If you have not done so already download: Dataset for Heterodyne Data Reduction/Analysis Tutorial 1 (Filesize: ~77MB) containing the raw data. Copy this file to a suitable directory to work in (~/raw is used in the following example):
cp JCMT_HETERODYNE_tutorial_2016.tar.gz ~/raw
Switch to the raw directory and open the tarball:
tar xvzf JCMT_HETERODYNE_tutorial_2016.tar.gz
This directory should now contain a number of .sdf files. To being the reduction process it is recommended moving to a new folder (you might like to call this reduced). In this folder you should set up ORAC-DR for ACSIS data processing, and specify that the reduced files will be written to the current working directory:
oracdr_acsis -cwd
By default, ORAC-DR will issue a warning that the default input data directory does not exist. It is therefore necessary to manually set the ORAC_DATA_IN environment variable.
In the Bash or zsh shell, type:
export ORAC_DATA_IN=.
Or, in the TC-shell or C-shell, type:
setenv ORAC_DATA_IN .
now create a file harp_files.list with the absolute file path to the raw data. This can be done via:
ls /absolute/file/path/a20070705_0003[8-9]*.sdf > harp_files.list
It should now be possible to run ORAC-DR on the raw HARP data by typing the following:
oracdr -files harp_files.list -nodisplay -log sf
To view the reduced data cube simply run GAIA:
gaia ga20070705_38_1_integ.sdf
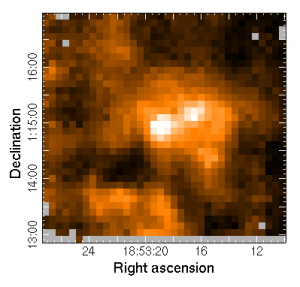
To view the integrated intensity image produced by the pipeline you can run GAIA:
gaia ga20070705_38_1_integ.sdf
STEP 2: Masking noise regions in the HARP CO (3-2) integrated intensity map
First we take the HARP CO (3-2) integrated intensity mask (ga20070705_38_1_integ.sdf) and remove the redundant third axis.
ndfcopy in=\"ga20070705_38_1_integ.sdf\(:,:\)\" out=ga20070705_38_1_integ_clip.sdf
we then remove noisy pixels/regions.
makesnr in=ga20070705_38_1_integ_clip.sdf out=ga20070705_38_1_integ_snr.sdf minvar=0
we then set the images to be bad or have a value of 1, depending if they are above or below a certain threshold in the signal-to-noise image. In this example we set this value to be 5*:
thresh in=ga20070705_38_1_integ_snr out=ga20070705_38_1_integ_snr_thresh thrhi=5 thrlo=5 newhi=1 newlo=bad
Now we apply this mask to the original data:
mult ga20070705_38_1_integ_snr_thresh.sdf ga20070705_38_1_integ.sdf out=ga20070705_38_1_integ_masked.sdf
NOTE: The above can also be done in a single step using an error clip to remove all pixels with a signal-to-noise-ratio greater than 5, however we will use the ga20070705_38_1_integ_snr_thresh file later. errclip in=ga20070705_38_1_integ_clip.sdf out=ga20070705_38_1_integ_masked.sdf limit=5 mode=snr
STEP 3: Convert the HARP integrated data from K to pW
To be converted to pW (the raw units of SCUBA-2 data) from K (the raw units of HARP data) we first need to correct the data for HARP efficiencies and transform the data from TA* to Tmb using ηmb = 0.64 from the HARP instrument page.
cdiv in=ga20070705_38_1_integ_masked out=ga20070705_38_1_integ_masked_tmb scalar=0.64
It is good practice to update the units and the data label as we progress. Note we can also track our progress using the kappa command hislist.
setunits ga20070705_38_1_integ_masked_tmb units=\"K km/s \"
setlabel ga20070705_38_1_integ_masked_tmb label=\"Tmb\"
We then need to calculate the conversion from K to pW, known as the C factor. This is explained more fully on the SCUBA-2 Molecular Line Contamination page. The C factor is known to depend on the amount of water in the atmosphere and so the initial steps for the conversion is to determine the PWV value/the conditions that the HARP data were obtained under. This is done using fitslist:
fitslist ga20070705_38_1_integ_masked_tmb.sdf | grep WVMTAU
The WVM opacity reported at the JCMT is obtained by the 183GHz WVM instrument installed inside the JCMT’s receiver cabin (the reported 186GHz in some observations is a bug/typo in our software – please ignore). The value reported, alhtough obtained at 183GHz, is reported in terms of Tau225GHz. The reason the WVM is reported at 225GHz is historical and relates to a time when the JCMT used the CSO tipping radiaometer (operating at 225GHz) for reference.
The conversion from Tau225GHz to PWV is given on the weather page and in Dempsey et al. 2013:
Tau225GHz = 0.04 PWV + 0.017
We can calculate this PWV value using kappa calc function, where WVM tau was recorded to be 0.108. We call the WVM tau in kappa calc: PT.
calc exp="(PT-0.017)/0.04" PT=0.108
From this we find the PWV to be 2.275.
Knowing the PWV value we can calculate the C850 factor for this data using the relations given on the SCUBA-2 contamination page. We can calculate this C850 factor using kappa calc function, where PA though PE are the C function coefficients at 850 microns and PW is the PWV value previously calculated.
calc exp="(PA)+(PB*PW)-(PC*PW**2)+(PD*PW**3)-(PE*PW**4)" PA=0.574 PB=0.1151 PC=0.0485 PD=0.0109 PE=0.000856 PW=2.275
we then apply this value to the data:
cmult in=ga20070705_38_1_integ_masked_tmb.sdf out=ga20070705_38_1_integ_masked_Cfactor scalar=0.69
Again it is good practice to update the units and the data label as we progress.
setunits ga20070705_38_1_integ_masked_Cfactor units=\"mJy/beam\"
setlabel ga20070705_38_1_integ_masked_Cfactor label=\"pseudo Flux Density\"
This gets the data into mJy/beam / (K/km/s). We will be applying this to the raw SCUBA-2 data so need this in pW, so we divide by the standard FCF (we chose the beam FCF as we applied the ηmb to the HARP data). Note the standard beam FCF is given in units of Jy/pW/beam (so we multiply by 1000 to get to mJy/pW/beam):
cdiv in=ga20070705_38_1_integ_masked_Cfactor.sdf out=ga20070705_38_1_integ_masked_CfactorpW.sdf scalar=537000.0
Updating the units (keeping the label as pseudo Flux Density):
setunits ga20070705_38_1_integ_masked_CfactorpW units=\"pW\"
We want to subtract this HARP data from the raw SCUBA-2 data. We will do this by inserting this data as a “fakemap”. Remember we are considering the CO (3-2) emission as contamination. As a result we invert the data:
cmult in=ga20070705_38_1_integ_masked_CfactorpW.sdf out=ga20070705_38_1_fakemap.sdf scalar=-1.0
setlabel ga20070705_38_1_fakemap label=\"inverted pseudo Flux Density\"
You now have produced the “fakemap” – ga20070705_38_1_fakemap.sdf – that will be required in step 5 of this process.
STEP 4: Creating SCUBA-2 850 micron emission reference map
If you have run the SCUBA-2 Data Reduction/Analysis Tutorial 1 and produced s20120501_00068_850_reduced.sdf you may skip this step but please rename the file to become: s20120501_00068_850_reference.sdf
If you have not done so already download: Dataset for SCUBA-2 Data Reduction/Analysis Tutorials 1 – 3 (Filesize: ~650 MB). Note that when you extract the data from this tarball the SCUBA-2 data will end up in a file named /tutorial/raw
tar xvzf JCMT_SCUBA-2_tutorial_2016.tar.gz
You are now advised to move to the directory where the other reduced HARP files are located. In this folder set up ORAC-DR for SCUBA-2 data processing, and specify that the reduced files will be written to the current working directory:
oracdr_scuba2_850
By default, ORAC-DR will issue a warning that the default input data directory does not exist. It is therefore necessary to manually set the ORAC_DATA_IN environment variable:
In the Bash or zsh shell, type:
export ORAC_DATA_IN=.
Or, in the TC-shell or C-shell, type:
setenv ORAC_DATA_IN .
now create a file scuba2_files.list with the absolute file path to the raw data.
ls /absolute/file/path/s8?20120501_00068*.sdf > scuba2_files.list
It should now be possible to run ORAC-DR on the raw SCUBA-2 data by typing the following:
oracdr -files scuba2_files.list -nodisplay -log sf
which will produce file – s20120501_00068_850_reduced.sdf. Rename that file to be s20120501_00068_850_reference.sdf :
mv s20120501_00068_850_reduced.sdf s20120501_00068_850_reference.sdf
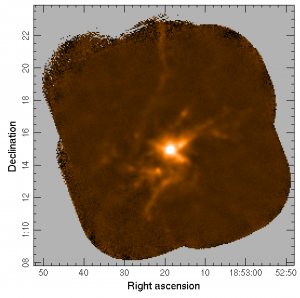
gaia s20120501_00068_850_reference.sdf
STEP 5: Creating SCUBA-2 with HARP CO subtracted from the 850 micron emission
We now wish to re-reduce the SCUBA-2 data using the HARP data as a fakemap. To do this we must ensure the HARP and SCUBA-2 data are aligned. To do this we first need to remove the third axis from the SCUBA-2 data. We can see this third axis if we do an ndftrace:
ndftrace s20120501_00068_850_reference.sdf
removing the third axis
ndfcopy in=\"s20120501_00068_850_reference.sdf\(:,:\)\" out=s20120501_00068_850_reference_clip.sdf
and now we can align the HARP data to the SCUBA-2 data.
wcsalign in=ga20070705_38_1_fakemap.sdf out=ga20070705_38_1_fakemap_aligned.sdf ref=s20120501_00068_850_reference_clip.sdf accept
Next we need to tell ORAC-DR pipeline that we wish to use this HARP data as a fake map. This is a three step process:
i) check the reduction recipe used during the original/reference SCUBA-2 reduction.
fitsval s20120501_00068_850_reference.sdf RECIPE
In this example we see that the map has been reduced using the REDUCE_SCAN recipe. Classically the REDUCE_SCAN recipe file calls the dimmconfig_jsa_generic.lis configuration file. We need to know this as we want to add a parameter to this configuration file.
ii) use your favorite editor to make a new configuration file: dimmconfig-contamination.lis. We will use this to point to the want this to point to the dimmconfig_jsa_generic.lis configuration file and include the newly made fakemap.
^$STARLINK_DIR/share/smurf/dimmconfig_jsa_generic.lis
fakemap=ga20070705_38_1_fakemap_aligned.sdf
iii) in order for this new dimmconfig file to be picked up during the reduction we will link it to a recipe parameter file. This file will be called myparams.ini this will need to contain the following:
[REDUCE_SCAN]MAKEMAP_CONFIG=dimmconfig-contamination.lis
and now run the reduction:
oracdr -files scuba2_files.list -recpars myparams.ini -nodisplay -log sf
which will produce a file called s20120501_00068_850_reduced.sdf. For clarity, rename that file to be s20120501_00068_850_corrected.sdf:
mv s20120501_00068_850_reduced.sdf s20120501_00068_850_corrected.sdf
This is now the SCUBA-2 850 micron emission with the CO (3-2) emission removed over a region of the map.
gaia s20120501_00068_850_corrected.sdf
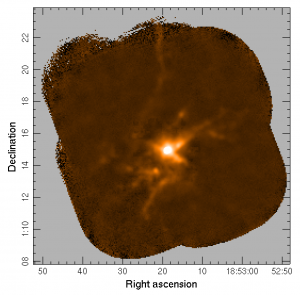
SCUBA-2 map of g34 with the emission from CO (3-2) as provided by HARP, removed from the raw SCUBA-2 data during the initial step of the reduction process.
STEP 6: Comparing SCUBA-2 reductions:
To create the contamination map we simply want to perform the following calculation:
(SCUBA-2 map – HARP CO emission) / SCUBA-2 map = the dust fraction
if HARP contributes 0 to the emission then we obtain a value 1 from the above equation indicating that 100% of the emission originates from continuum emission and not from emission at the CO frequency.
To perform this on the command line we use:
div s20120501_00068_850_corrected.sdf s20120501_00068_850_reference.sdf div.sdf
again we remove the redundant third axis:
ndfcopy in=\"div.sdf\(:,:\)\" out=dust_fraction.sdf
We only want to consider the above map where we know we have associated HARP data. We should also be careful not to include the edges of the HARP map as they may suffer from issues resulting from a step function at the boundary in the reduction.
To trim the edges we first inspect the original HARP map size:
ndftrace ga20070705_38_1_integ_snr_thresh.sdf
To trim the edges of the map by five times the SCUBA-2 beam we will need to clip the map edges by 10 pixels*
ndfcopy in=\"ga20070705_38_1_integ_snr_thresh.sdf\(-7:8,-6:7\)\" out=ga20070705_38_1_integ_snr_thresh_clipped
again we ensure the maps are aligned:
wcsalign in=ga20070705_38_1_integ_snr_thresh_clipped.sdf out=ga20070705_38_1_integ_snr_thresh_clipped_aligned ref=dust_fraction.sdf accept
and then extract only the region of interest
mult ga20070705_38_1_integ_snr_thresh_clipped_aligned.sdf dust_fraction.sdf out=dust_fraction_cropped.sdf
and remove excess bad pixels:
ndfcopy in=dust_fraction_cropped.sdf out=dust_fraction_clipped.sdf trimbad=true
It is then recommended to inspect this final map, using the stats, histat and histogram commands. Using
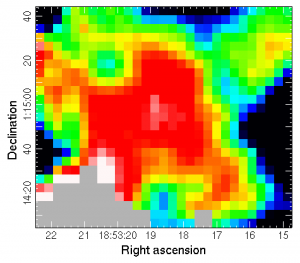
gaia dust_fraction_clipped.sdf
we see the following dust fraction map.
*This is by no means a comprehensive guide and you should consider your own data/cuts to data before making decisions on where/exactly how to inspect the resulting contamination map.